POInTDownload Help
Topics
Overview
POInTdownload is the batch interface to the POInT datasets, allowing users to download sets of gene trees and coding sequences corresponding to POInT pillars that meet certain criteria. For instance, one could download all high-confidence pillars which retain duplicated genes in all genomes, or download known single-copy orthologs with similar high confidence. Results are returned as UNIX tarfiles, including gene trees in the selected format as well as the corresponding DNA CDS sequences in FASTA format.
Quick-Start Guide
Example POInTdownload window
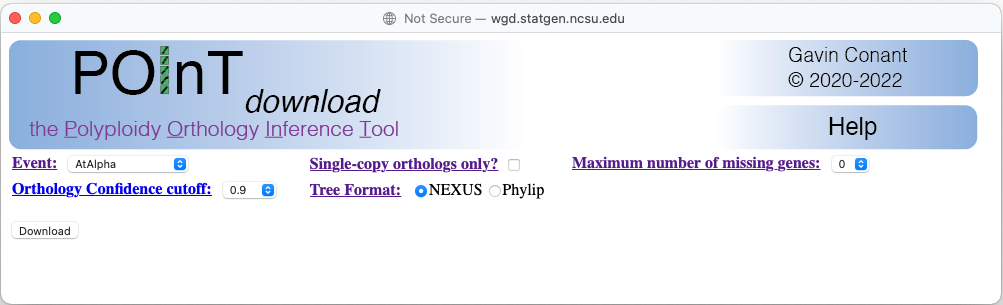

Maximum number of missing homoeologs
Gives the maximum number of species from which a surviving duplicate may be missing. Hence a value of 0 means that no single copy genes are allowed. The largest value is set by the number of genomes in the POInT dataset for that event, which, for a WGD, would correspond to allowing all possible combinations of duplicated and single-copy genes.
Subgenomes
For polyploidy events with evidence for signficant biased fractionation (currently all events save that in yeast), POInT can assign orthologous single-copy genes to subgenomes, which are inferred to derive from one of the two allopolyploid parents. In the case of single-copy orthologys, the confidence in this assignment is the same as the overall "pillar" confidence. Hence, when single-copy orthologs are requested from the download engine, you have the option of requesting all such orthologs, or only those deriving from the subgenome that is least fractionated (has the most surviving genes) or the most fractionated subgenome (fewest surviving genes). For the hexaploids, there is also a subgenome of intermediate fractionation.
Orthology confidence cutoff
POInT's orthology inferences have associated statistical confidence representing the likelihood of the best orthology assignment over the sum of the likelihoods of all possible orthology assignments. Hence higher values give greater confidence in the orthology assignments returned.
These confidences include assigning each gene to a subgenome, and confidence values range from 0.0-1.0. For events without biased fractionation (here the Yeast WGD), the orthology assignments are degenerate with respect to subgenomes, meaning that the assignment 000..0-111..1 and 111..1-000..0 are equivilent. As such, while the potential confidence values range from 0.0-1.0 as usual, those confidence values do not assign genes to subgenomes and the picklist for subgenomes when orthologs are requested is not populated.
Single-copy orthologs only
This selection causes the download to include only sets of single-copy genes where the gene in every species is inferred to be the ortholog of that in every other species at the confidence selected. Hence, this option will change the screen to a new format:
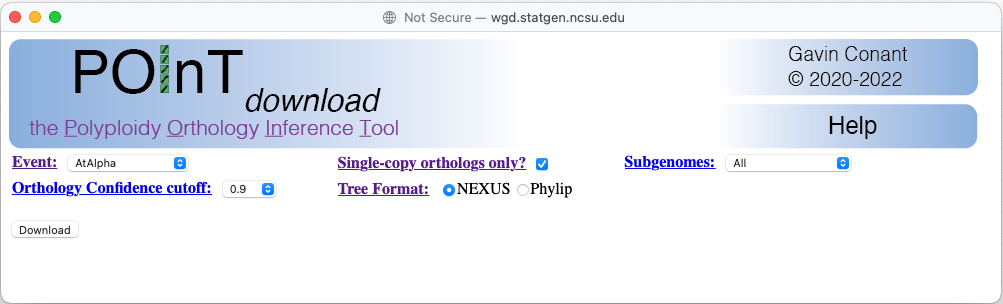
In particular, the picklist for selecting the number of surviving homoeologs disappears and is replaced with a picklist allowing you to select which subgenome the orthologs derive from.
Download
Once you have selected the desired gene sets, this button performs the download.
Help
Opens this help page.

